Trey Ideker Laboratory
The past decade has seen an explosion in “genome-era” technologies which profile genes, proteins, metabolites and the intricate web of interconnections among them. Equally important is the development of methods to assemble these diverse measurements into models of functional networks and pathways within the cell. These pathway maps will provide essential information as doctors struggle to interpret the flood of genetic and clinical data that can now be collected for a patient.
We are working in several areas that we believe will be critical for assembling network models and for using them in a clinical setting:
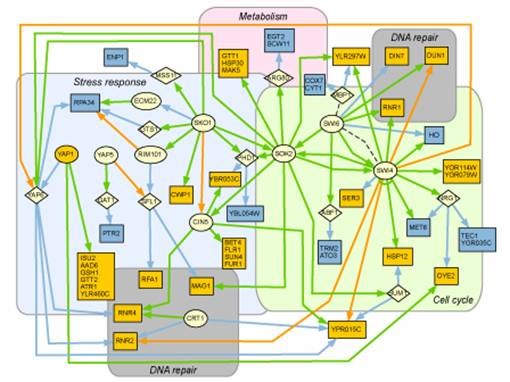
- Mapping the genetic network underlying the response to DNA damage. Failure of cells to respond to DNA damage is a primary step in the onset of cancer and is a key mechanism of environmental toxicity. Consequently, cells have evolved complex repair and stress responses that are highly conserved across the eukaryotic kingdom, from yeast to humans. We will describe our ongoing efforts to apply ChIP-sequencing and synthetic-lethal screens to map how the cell's transcriptional network is remodeled by DNA-damaging conditions.
- Network-based biomarkers for disease diagnosis and personalized medicine. Genetic biomarkers are typically thought of as individual genes and proteins— for example using prostate specific antigen (PSA) as a marker for prostate cancer. Recently, we have shown that networks can also serve as powerful biomarkers and in many cases are more predictive than any individual gene. Our approach is to project gene and/or protein expression profiles of each patient onto the known human genetic network map to identify pathways that are predictive of disease. This “network-based” biomarker approach has shown improved accuracy in diagnosis of breast and lung cancer as well as NF-kB activation state.
- Protein network comparative genomics. We are developing a library of standard approaches for comparing protein interaction networks across species, conditions, and network types. We will describe network comparisons to study the protein interaction network of Plasmodium, the pathogenic protozoan that causes malaria, which surprisingly is quite divergent from other known networks. We are also working with Dr. Sumit Chanda at the Burnham Institute to identify protein networks essential for HIV infection and how these differ from RNA and DNA viruses.
For professional distribution of our network-based technologies, we are developers of the Cytoscape platform, an Open-Source software environment for visualization and analysis of biological networks and models.
Israni Biomedical Research Facility, Office 3A13
3147 Biomedical Sciences Way, La Jolla, CA 92093
